Workflow optimization
Pathology departments in Western Norway are processing over 500 000 slides. We need to optimize our work processes to report a diagnosis as fast as possible.
Work Package 7: Workflow optimization
Motivation & Background
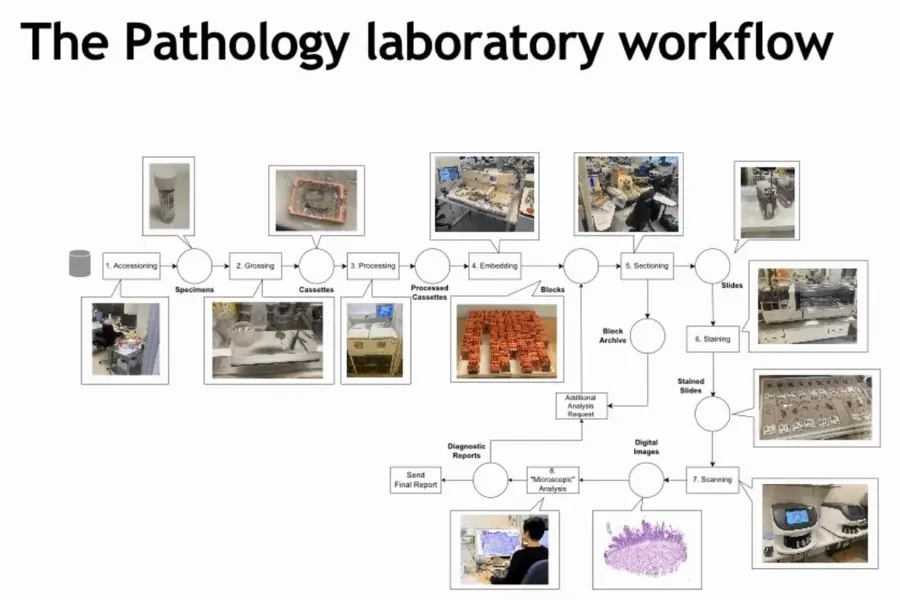
Pathology literally means the study of diseases. Within the complex organization of a hospital, the pathology department plays an important role as a diagnostic component. It receives tissue and cell specimens from patients, which are analyzed under a microscope where a pathologist writes a diagnostic report. The latter decides the further treatment of a patient. However, before a specimen can be analyzed under the microscope, it must undergo an elaborate preparation process in the laboratory:
The above animation in figure 1 depicts the various stations, which a tissue specimen (this branch of pathology is known as histology) passes through during the laboratory preparation process. Several of these activities are manual activities and require trained laboratory personnel (lab technicians and pathology residents). As with many resources in the healthcare sector, the availability of laboratory workers is scarce. Simultaneously, pathology departments are experiencing a steady increase in the number of incoming specimens. As a result, the turnaround times, i.e., the time patients have to wait in between sending their specimen and receiving a diagnostic response, have increased over the last years.
The main objective of this work package is to investigate how the turnaround times of the pathology department in Bergen may be reduced by integrating modern workflow management and observability solutions (the keyword here is “digital twin”) as well as means of optimal resource allocation. Concretely, we are doing research on the following topics:
1. Event Data and Process Mining
2. Statistics and Process Modelling
3. Simulation and Prediction
4. Optimization and Self-Adapting Systems
1. Event Data and Process Mining
The foundation of our approach is based on something called an event log. Abstractly, the latter is defined as a set of cases, where a case is simply an ordered sequence of events. Such event logs provide insightful information about the execution of processes a.k.a. workflows.
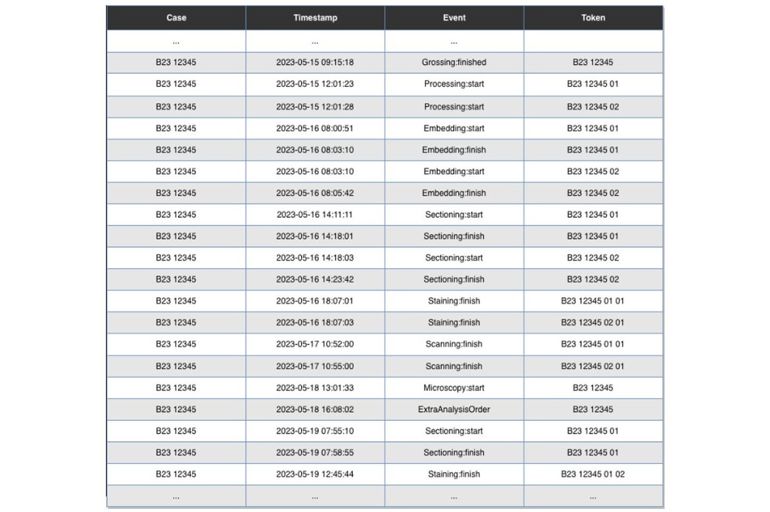
Figure 2 depicts how such an event log may look like for our “Histo-Pathology”-workflow. The research discipline that investigates event logs and turns them into valuable knowledge is called process mining, which lies in the intersection between data science and process science. The objective here is to obtain a process model from the raw event logs. A process model is an abstract yet formal representation of a real-world process and comprises:
- a control-flow perspective (i.e., what events and activities exist in a process and how is there causal ordering?),
- an organizational perspective (i.e., what resources and actors are involved and how are they related?),
- a data-perspective (i.e., what elements flow through the process, what information do they contain, and what rules do they obey?),
- a time-perspective (i.e., how long do certain activities take?)
The PiV project combines domain-specific knowledge and state-of-the-art process mining research knowledge in order to extract a process model of pathology from database of the laboratory information system, which is used by the department in Bergen.
2. Statistics and Process Modelling
After having utilized the raw event data in the form of a process model, one can start to analyze and diagnose the process. This can mean to compute simple statistics and key performance indicators (e.g., number of answered cases today, sectioned blocks, …) and to present them in a useful way. This enables us to answer the question about “What is happening” and start to investigate “why is something happening”. The results of this phase are dashboards and solutions for statistical analysis of the underlying process data set.
3. Simulation and Prediction
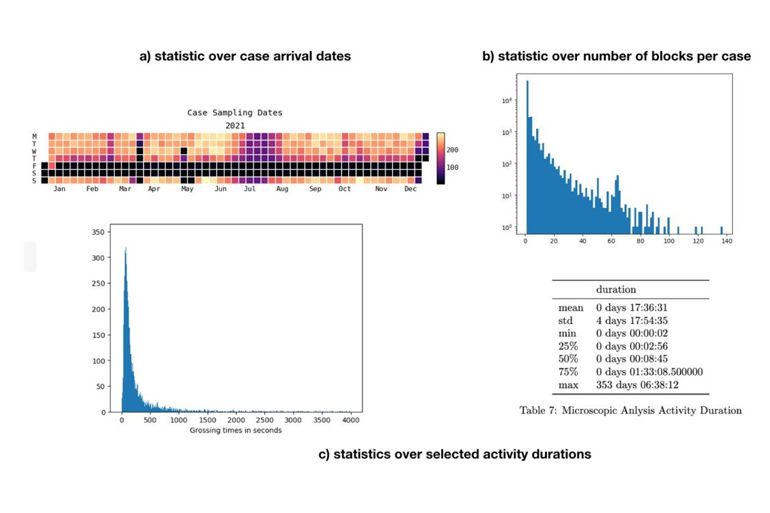
With the availability of a formal process model and statistical models about resource and time resources, we are able to run discrete event simulations (DES). Those simulations can be run with various configurations and parameters and therefore allow to “play around” with the process explore how different factors affect the process and turn-around-times. The latter allows to answer the question of “What will happen” and to predict future workloads and process performance.
4. Optimization and Self-Adapting Systems
The final stage of the project is about combining all the artefacts produces by the aforementioned stages in order to create a system, which produces suggestions for possible interventions that will reduce turn-around-times, e.g., by assigning the scarce worker resources in such a way that an optimal utilization is achieved or by dynamically changing priorities of individual process items. There are several scientific disciplines that can help us in finding answers here: Automated Planning and Reinforcement Learning are sub-fields of Artificial Intelligence where an agent learns to solve problems on its own and continuously improves its performance. Mathematical Optimization is an important part of Operations Research, which seeks to find an optimal assignment of parameter values to reduce an objective cost function. Our general objective of reducing the turn-around-time of the Pathology laboratory is actually an instance of the resource-constrained project scheduling (RCPS) problem, which is a special case of an optimization problem.
Contact and Collaboration
If you think that one or more of the topics above sound interesting, you are more than welcome to contact us! We are always looking for potential collaborators. It is also possible to write Bachelor and Master theses with us if you are a student.
The main contact person for WP7 is Associate Professor Patrick Stünkel.
Software and other technical artefacts can be found on our GitHub page.